Anatomical and Lesion Segmentation
Location-Aware Parameter Fine-Tuning for Multimodal Image Segmentation | Link in here
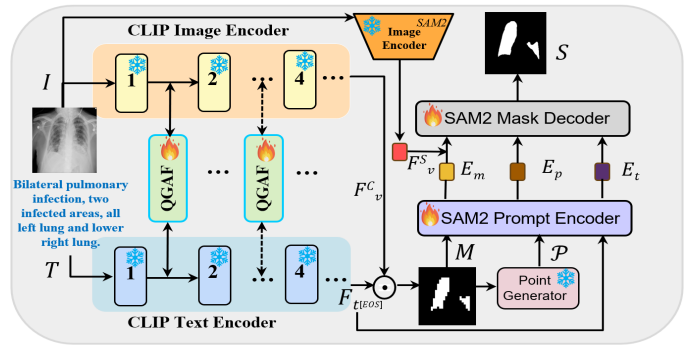
Accurate segmentation of lung infection regions is critical for early diagnosis and quantitative assessment of disease severity. However, existing segmentation methods largely depend on high-quality, manually annotated data. Although some approaches have attempted to alleviate the reliance on detailed annotations by leveraging radiology reports, their complex model architectures often hinder practical training and widespread clinical deployment. With the advent of large-scale pretrained foundation models, efficient and lightweight segmentation frameworks have become feasible. In this work, we propose a novel segmentation framework that utilizes CLIP to generate multimodal high-quality prompts, including coarse mask, point, and text prompts, which are subsequently fed into the Segment Anything Model 2 (SAM2) to produce the final segmentation results. To fully exploit the informative content of medical reports, we introduce a localization loss that extracts positional cues from the text to guide the model in localizing potential lesion regions.
Prototype-Based Image Prompting for Weakly Supervised Histopathological Image Segmentation | Link in here
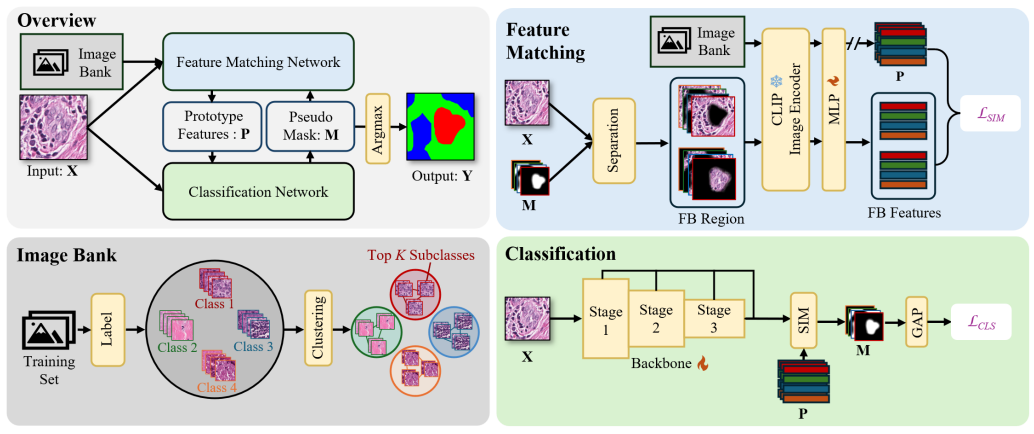
Weakly supervised image segmentation with image-level labels has drawn attention due to the high cost of pixel-level annotations. Traditional methods using Class Activation Maps (CAMs) often highlight only the most discriminative regions, leading to incomplete masks. Recent approaches that introduce textual information struggle with histopathological images due to inter-class homogeneity and intra-class heterogeneity. In this paper, we propose a prototype-based image prompting framework for histopathological image segmentation. It constructs an image bank from the training set using clustering, extracting multiple prototype features per class to capture intra-class heterogeneity. By designing a matching loss between input features and class-specific prototypes using contrastive learning, our method addresses inter-class homogeneity and guides the model to generate more accurate CAMs.
Multi-Degradation-Adaptation Network for Fundus Image Enhancement with Degradation Representation Learning | Link in here
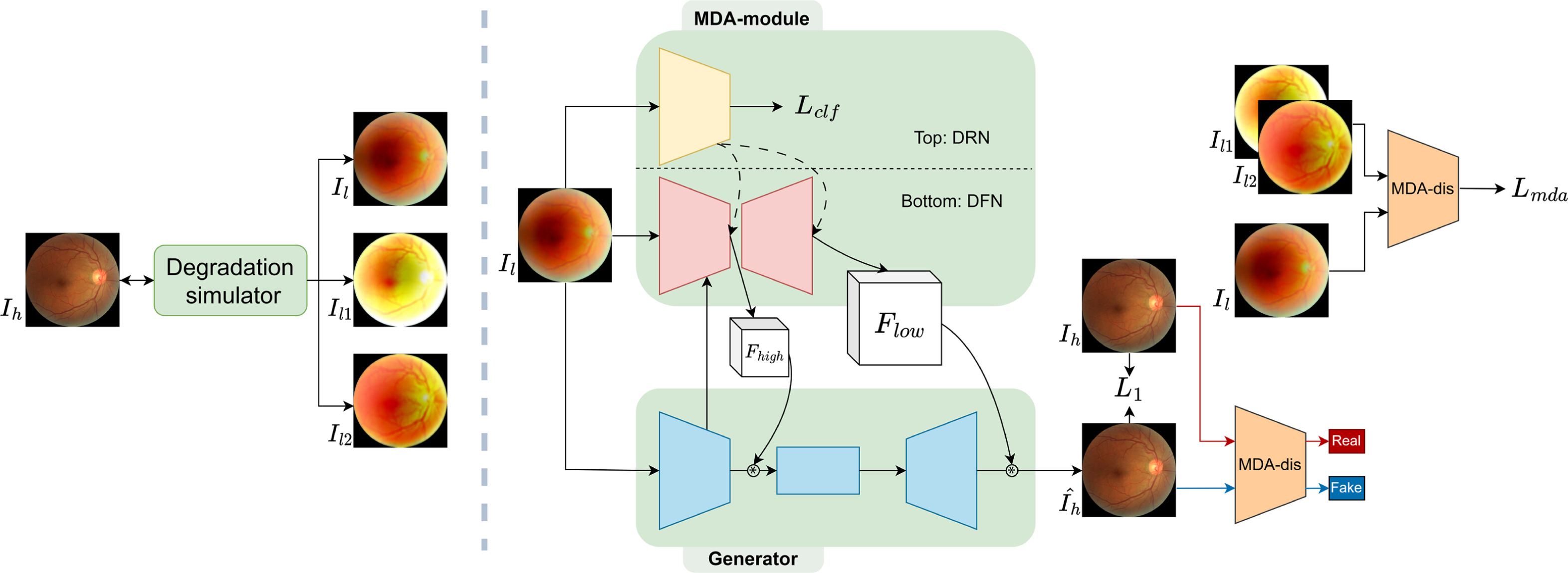
Fundus image quality serves a crucial asset for medical diagnosis and applications. However, such images often suffer degradation during image acquisition where multiple types of degradation can occur in each image. Although recent deep learning based methods have shown promising results in image enhancement, they tend to focus on restoring one aspect of degradation and lack generalisability to multiple modes of degradation. We propose an adaptive image enhancement network that can simultaneously handle a mixture of different degradations. The main contribution of this work is to introduce our Multi-Degradation-Adaptive module which dynamically generates filters for different types of degradation. Moreover, we explore degradation representation learning and propose the degradation representation network and Multi-Degradation-Adaptive discriminator for our accompanying image enhancement network.
SAC-Net: Learning with weak and noisy labels in histopathology image segmentation | Link in here
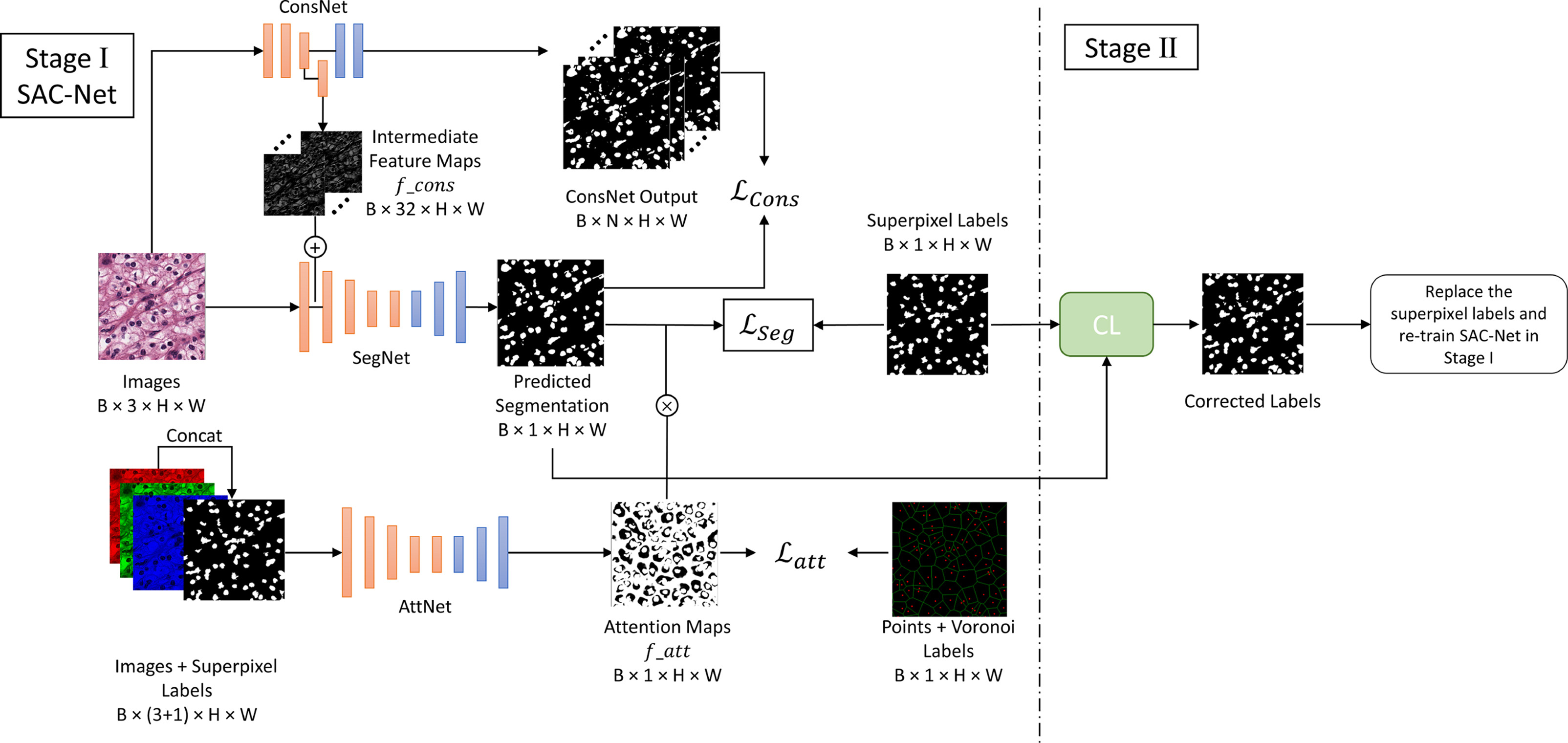
Deep convolutional neural networks have been highly effective in segmentation tasks. However, segmentation becomes more difficult when training images include many complex instances to segment, such as the task of nuclei segmentation in histopathology images. Weakly supervised learning can reduce the need for large-scale, high-quality ground truth annotations by involving non-expert annotators or algorithms to generate supervision information for segmentation. However, there is still a significant performance gap between weakly supervised learning and fully supervised learning approaches. In this work, we propose a weakly-supervised nuclei segmentation method in a two-stage training manner that only requires annotation of the nuclear centroids. First, we generate boundary and superpixel-based masks as pseudo ground truth labels to train our SAC-Net, which is a segmentation network enhanced by a constraint network and an attention network to effectively address the problems caused by noisy labels. Then, we refine the pseudo labels at the pixel level based on Confident Learning to train the network again.
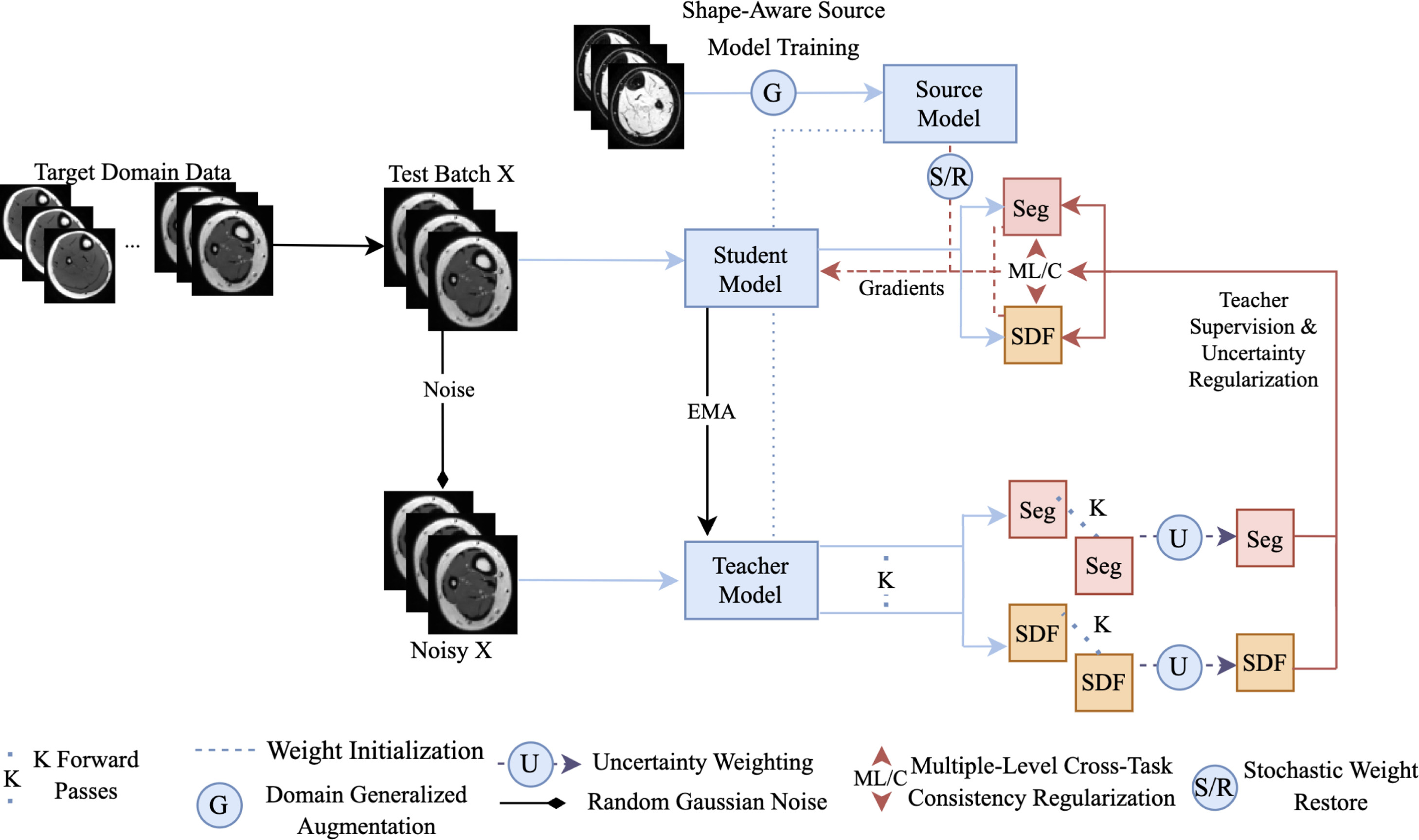
Improving cross-domain generalizability of medical image segmentation using uncertainty and shape-aware continual test-time domain adaptation | Link in here
Empirical risk minimization (ERM)-trained models often fail to perform adequately on a severely drifted target domain, resulting in unsatisfactory adaptation results. To tackle this issue, we propose a generalizable continual test-time adaptation (CTTA) framework. First, we incorporate domain-invariant shape modeling into the model and train it using domain-generalization (DG) techniques, promoting target-domain adaptability regardless of the severity of the domain shift. Then, an uncertainty and shape-aware mean teacher network performs adaptation with uncertainty-weighted pseudo-labels and shape information. As part of this process, a novel uncertainty-ranked cross-task regularization scheme is proposed to impose consistency between segmentation maps and their corresponding shape representations, both produced by the student model, at the patch and global levels to enhance performance further. Lastly, small portions of the model’s weights are stochastically reset to the initial domain-generalized state at each adaptation step, preventing the model from ‘diving too deep’ into any specific test samples.