Cancer Analysis
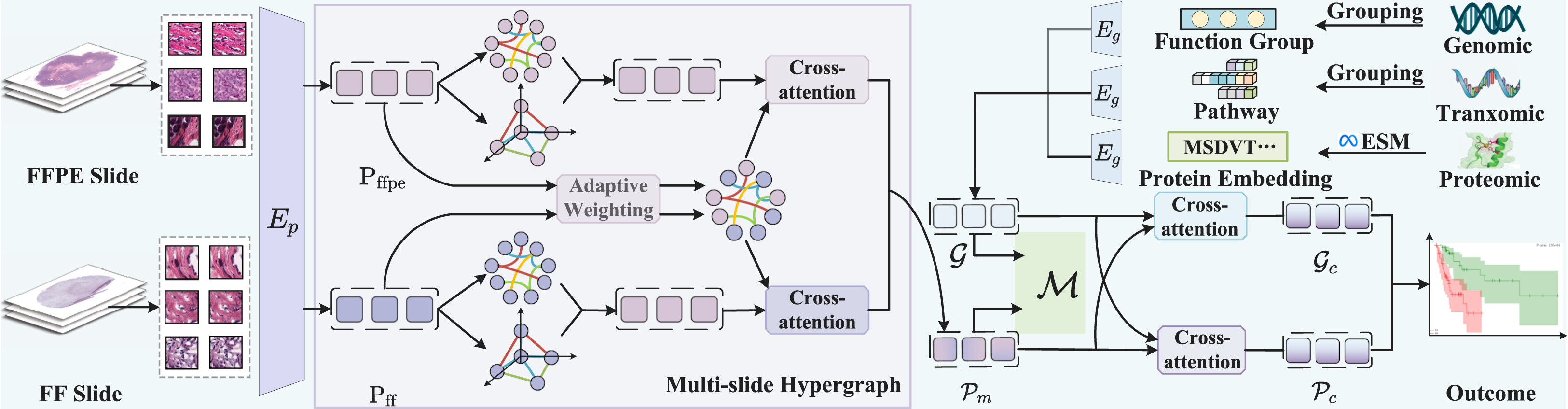
M3Surv: Fusing Multi-Slide and Multi-Omics for Memory-Augmented Robust Survival Prediction | Link in here
Existing multimodal survival prediction methods typically integrate only Formalin-Fixed Paraffin-Embedded (FFPE) slides with a single omics type, such as genomics, overlooking Fresh Frozen (FF) slides that better preserve molecular information, as well as richer multi-omics data like proteomics and transcriptomics. More critically, the complete absence of certain modalities due to clinical constraints (e.g., time or cost) severely limits the applicability of conventional fusion models that rely on inter-modality correlations. To address these gaps, we propose M3Surv, a framework designed to integrate multi-pathology slides (both FF and FFPE) with multi-omics profiles. For multi-slide fusion, we design a divide-and-conquer hypergraph learning approach to capture both intra-slide higher-order cellular structures and inter-slide relationships, yielding a unified pathology representation. To enrich the biological context, we integrate multi-omics data and employ interactive cross-attention to fuse the pathological and omics modalities. To tackle the missing modality, we introduce a prototype-based memory bank. During training, this memory bank learns and stores representative pathology-omics feature prototypes.
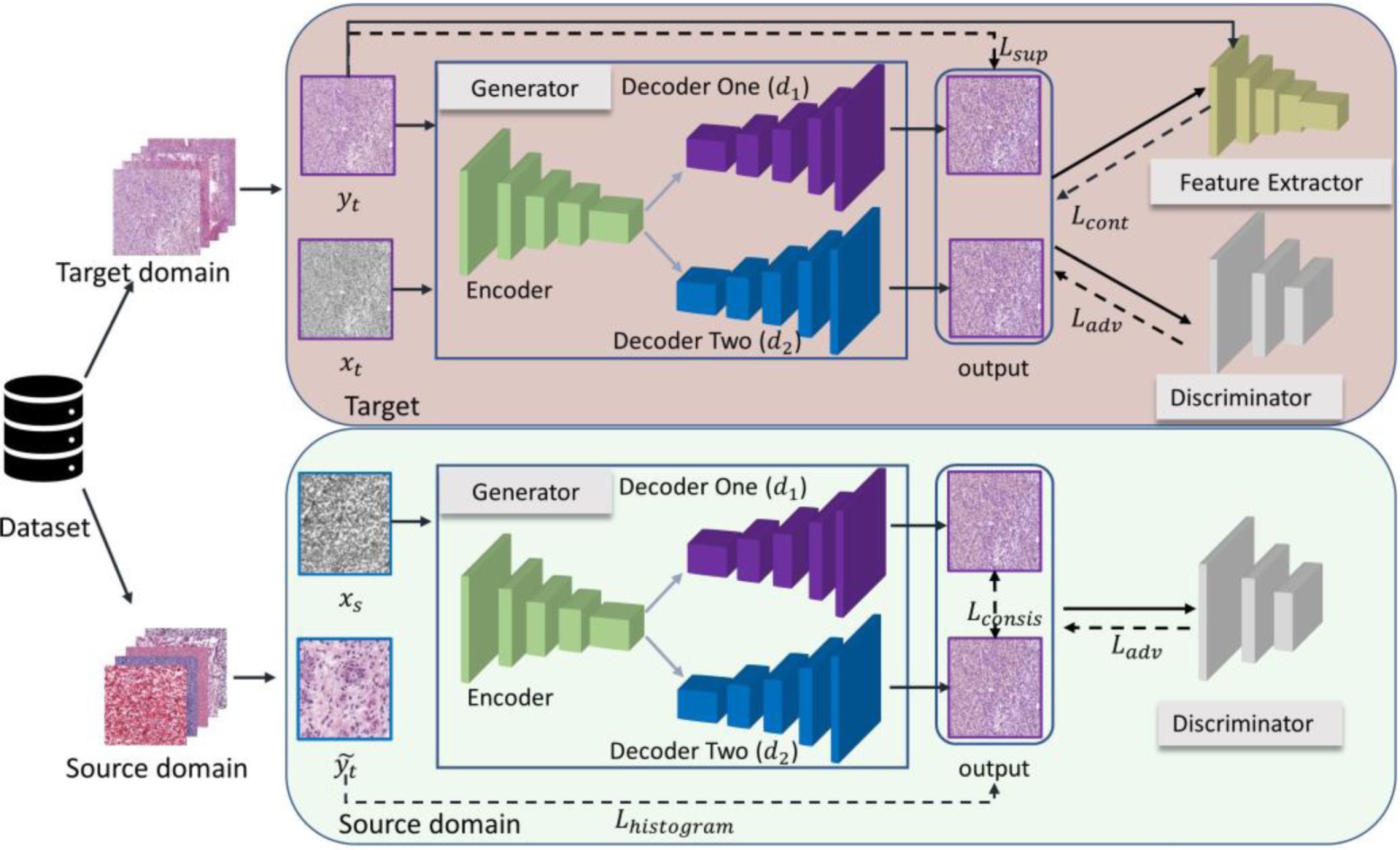
Colour Adaptive Generative Networks for Stain Normalisation of Histopathology Images | Link in here
Although deep learning-based stain normalisation methods have been proposed for histopathology image analysis, these methods are either trained fully with paired images from the target domain (supervised) or with unpaired images (unsupervised), suffering from either large discrepancy between domains or risks of undertrained/overfitted models when only the target domain images are used for training. We introduce a colour adaptive generative network (CAGAN) for stain normalisation which combines both supervised learning from target domain and unsupervised learning from source domain. Specifically, we propose a dual-decoder generator and force consistency between their outputs thus introducing extra supervision which benefits from extra training with source domain images. Moreover, our model is immutable to stain colour variations due to the use of stain colour augmentation. We further implement histogram loss to ensure the processed images are coloured with the target domain colours regardless of their content differences.
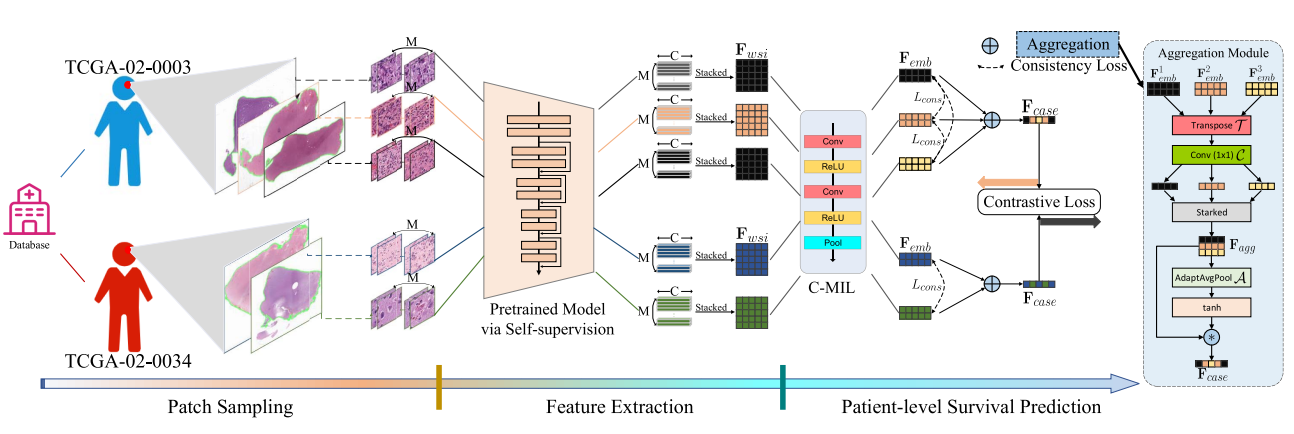
Cancer Survival Prediction from Whole Slide Images with Self-Supervised Learning and Slide Consistency | Link in here
Histopathological Whole Slide Images (WSIs) at giga-pixel resolution are the gold standard for cancer analysis and prognosis. Due to the scarcity of pixel- or patch-level annotations of WSIs, many existing methods attempt to predict survival outcomes based on a three-stage strategy that includes patch selection, patch-level feature extraction and aggregation. However, the patch features are usually extracted by using truncated models (e.g. ResNet) pretrained on ImageNet without fine-tuning on WSI tasks, and the aggregation stage does not consider the many-to-one relationship between multiple WSIs and the patient. In this paper, we propose a novel survival prediction framework that consists of patch sampling, feature extraction and patient-level survival prediction. Specifically, we employ two kinds of self-supervised learning methods, i.e. colorization and cross-channel, as pretext tasks to train convnet-based models that are tailored for extracting features from WSIs. Then, at the patient-level survival prediction we explicitly aggregate features from multiple WSIs, using consistency and contrastive losses to normalize slide-level features at the patient level.